ADME SARfari: A tool for predicting and comparing cross-species ADME targets
ADME
studies are focused on understanding the disposition of a compound
within an organism and the results of such studies play a critical role
in the drug development process. ADME studies (more commonly referred to
as pharmacokinetic or PK studies) are focused on 4 main areas:
Absorption, Distribution, Metabolism and Excretion. More information on
the PK measurement types can be found here.
Comparisons
of PK data across species is a potential problem drug researchers need
to deal with, as model organism studies are the primary source of such
data. For example, in an animal model study, which may be carried out on a compound as it passes through the drug development pipeline, is it
meaningful to compare clearance or bioavailability data from a mouse or
rat to human? Clearly there are many differences (physical, metabolic,
genetic,..), which make answering these types of questions difficult. Building tools which guide researchers to potential answers or provide a better understanding of the inter-species differences are of great value - leading us nicely to the focus of this blog post.
It turns out the ChEMBL database has a wealth of PK measurements data, which allows users to start asking ADME focused questions. You can access all of this data via the ChEMBL Interface or from one of our downloads, but in order to answer some of the more complex questions a significant amount of data processing first needs to take place. So to help the ChEMBL community get started analysing this data, we, in collaboration with our colleagues at GSK, set about building a new ADME focused system. The new system is called ADME SARfari and it aims to centralise all ADME data currently stored in ChEMBL and other related databases, as well as providing new tools to help interrogate the data.
In order to build the system we have pulled data from a number of sources, which include:
- ChEMBL - Bioactivity data, PK data and molecules.
- PharmaADME - An online resource providing a list ADME related human genes. We used this to build our primary list of ADME Targets, but have also added a couple of extra ones.
- The Human Protein Atlas - Protein expression data for human ADME Targets.
- ENSEMBL - Orthologue (using the Compara Service) and SNP data for ADME Targets.
- The Göttingen minipig and beagle dog genome predicted ADME Targets. GSK have sequenced the genomes of these two pharmaceutically significant species. More details on the genomes can be found here.
When you visit the site you will find it is divided into the following 7 sections:
- Home - Allows user to initiate a compound or protein focused search.
- Orthologues - A table of ADME orthologues. The first column of the table corresponds to the human ADME targets and additional columns correspond to targets found in model organisms.
- Tissues - Protein expression data for human ADME targets.
- Bioactivities - Bioactivity data and PK measurements for all ADME related targets in the system, which are also found in the ChEMBL database.
- Molecules - Distinct set of ChEMBL molecules linked to ADME related targets via the bioactivity data.
- Pharmacokinetic - A cross species comparison of PK data for compounds found in the ChEMBL database (see red heatmap image at top).
- About - More details on how to use the system
By clicking on the links above, you can view and download all of
the data associated within each of the sections. One exception is the
Pharmacokinetics section, as there is too much data to display in the
heatmap by default, so you must first initiate a search (we will
come back to this later). Now that you have a better idea of what is in
the system, what are the types of questions you can ask? Looking at the
homepage you will see there are 2 ways to initiate a search of the
system: Compound-initiated, using the chemical sketcher box and protein-initiated, using the BLAST or keyword search.
Protein Search (Using BLAST)
To run a BLAST search paste a sequence in the text box to the right on the homepage:
BLAST
search results will be displayed on the Orthologues page and only rows
which contain hits to your search query will be returned:
Clicking on the Tissues tab displays the protein expression levels of the human targets returned by the BLAST search:
Clicking on the Bioactivities tab returns all of the ChEMBL bioactivity data for targets returned by BLAST search:
Clicking on the Molecules tab returns the distinct set of compounds currently displayed in the Bioactivities section:
Clicking
on the Pharmacokinetics tab will provide a cross-species overview of
the PK data in ChEMBL for the compounds currently displayed in the
Molecules section (Note that not all compounds will have PK
measurements):
The following points help explain what the heatmap above is displaying:
- Each narrow row corresponds to a compound.
- Each column corresponds to a PK measurement in a specific organism.
- The PK measurements summarised in this view are Clearance (Cl), Cmax, Bioavailability (f), T1/2, Tmax and Volume of Distribution (Vd).
- The colour of each cell corresponds to a low, medium and high binned PK measurement, making it easier to compare values across species.
- The columns can be sorted by clicking on the header (see the second column, which corresponds to the Human Clearance data).
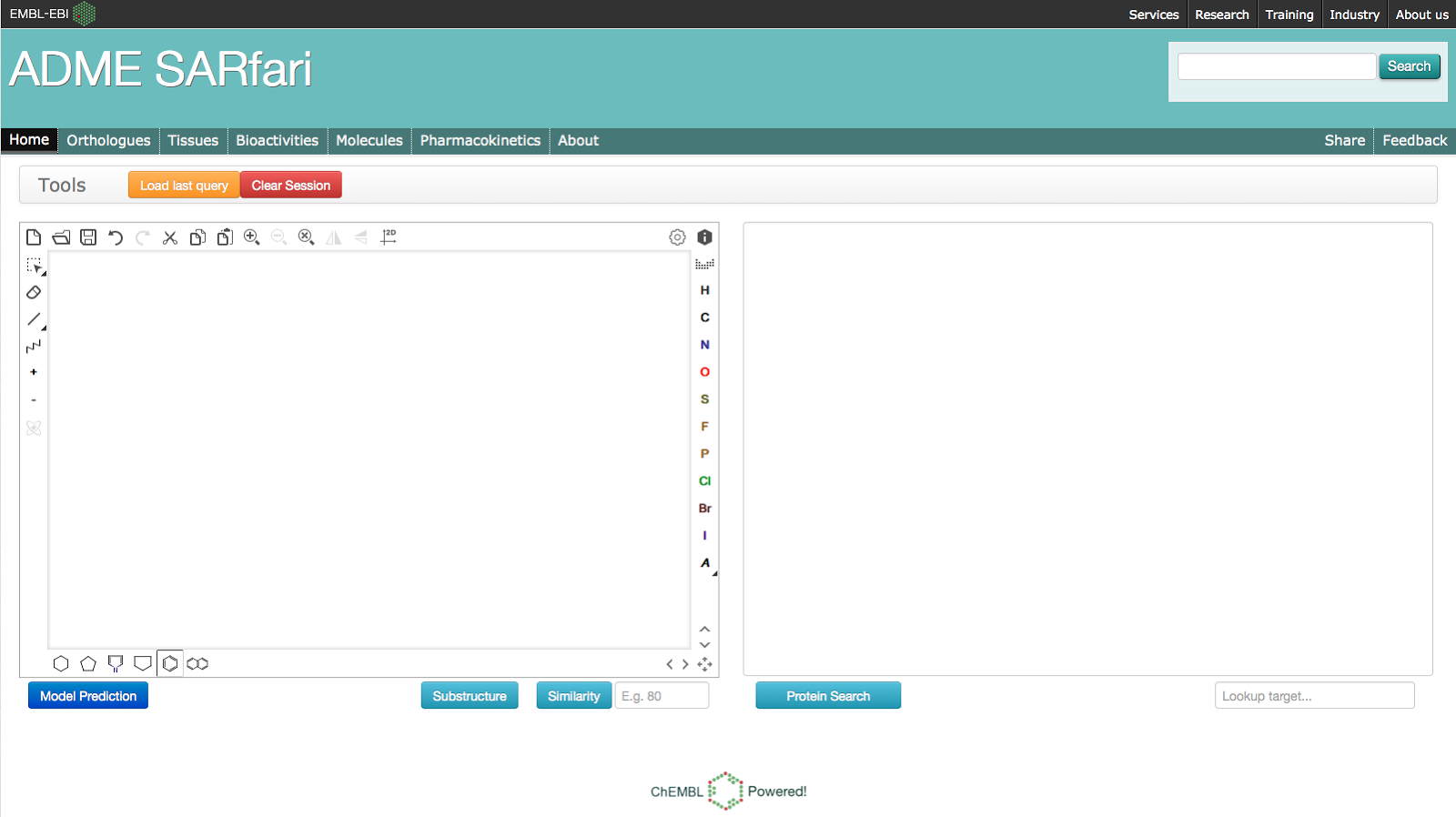
Compound Search
It is also
possible to search the system using a compound structure. Simply draw or
paste a structure into the compound sketcher on the left of the
homepage:
You
can then choose to run a substructure and similarity search. When you
run the search you will first be taken Molecules section and you can
then explore the other sections, which are all connected based on this
initial set of compounds.
Predictive Model Search
The
system also allows a user to predict which ADME protein target a
molecule will interact with. The binding data (displayed in the
Bioactivities section) has been used to build a multi-category naive Bayesian classifier. We will follow up with a more detailed post describing the
model building process, but essentially the model is able to predict if a
user-submitted molecule will interact with 133 of the ADME targets included
in the system. The About page provides some additional data on the targets included in the model.
To
run a search against the predictive model, draw or paste a structure
into the compound sketcher on the left of the homepage and hit the
"Model Prediction" button. You will be taken to the Orthologues section,
but only the rows which include a target predicted to interact with
submitted molecule (coloured green) will be on display:
We hope you find the ADME SARfari system useful and if you have any questions please let us know.
The ChEMBL Team