Paper: Chemical, Target, and Bioactive Properties of Allosteric Modulation
We have just had a paper accepted in PLoS Computational Biology on the work we've done on allosteric modulators (first mentioned on the blog here). The work is based on the mining of allosteric bioactivity points from ChEMBL_14. The data set of allosteric and non-allosteric interactions is available on our FTP site (here). This blogpost will just highlight some sections of the paper, but we would like to refer the interested reader to the full paper (here).
Dataset
The dataset contains ChEMBL annotated and cleaned data divided in both an 'allosteric' set and a 'non-allosteric' (or background) set. Abstracts and titles mentioning allosteric keywords were pulled and from the resulting papers we extracted the primary target and all bioactivities on this primary target. From the remainder of the papers we also retrieved the primary target and all bioactivities on this primary target in a similar manner.
Targets
When we observed the target distribution in both sets, we saw differences (see below ; also touched upon in the previous post). Targets that are known to be amenable to allosteric modulation are indeed well represented in our allosteric set (e.g. Class C GPCRs). However there are also some interesting observations that we did not expect (please see the paper for further details).
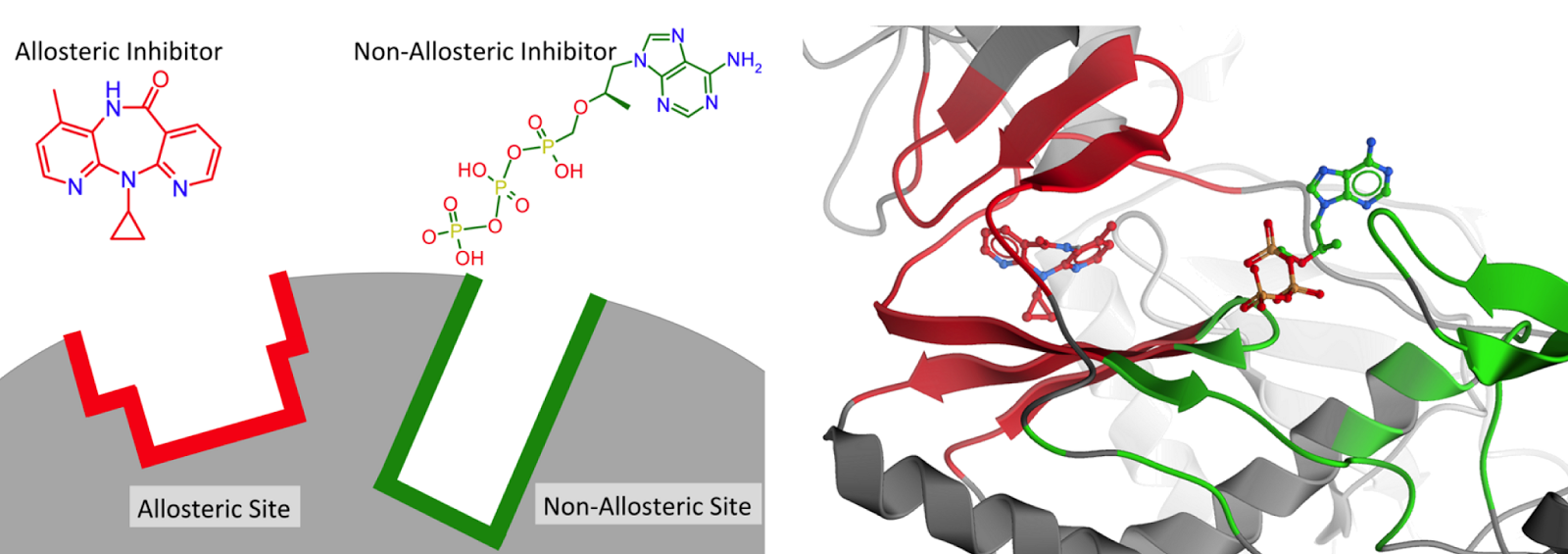
Chemistry
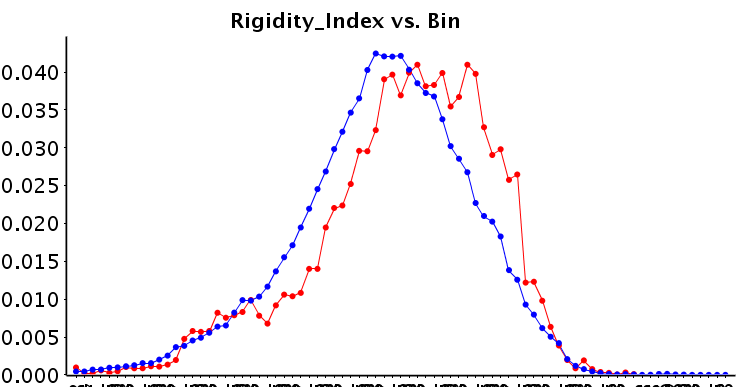
Obviously, as we are the ChEMBL group, we are interested in potential chemical differences between the allosteric and background set. Interestingly, the allosteric modulators appear to form a subset of the background set, rather than that they are distinct from the background set. We have calculated a large number of descriptors and compared the sets (median values, but also histograms; all available on the FTP). We observe that allosteric modulator molecules tend to be smaller, more lipophilic and more rigid. Although there is understandably a large variance over the diverse targets included in the set. Shown here is the rigidity index calculated over the full sets (L0), but when the target selection becomes more concise, the differences become more distinct.
Bioactivity
Likewise we observe differences between our allosteric subset and the background set with regard to bioactivity. While 'allosteric modulation' is a very diverse concept, in which the specific manner wherein the protein is influenced by the small molecule differs per protein - ligand pair, we do observe some general differences. From our data it appears that allosteric modulators bind with a lower affinity (on average) but similar ligand efficiency (on average) when compared to our background set. In the paper we provide a more extensive discussion on this observation and we would again refer the reader given the limited space here.
Classification models
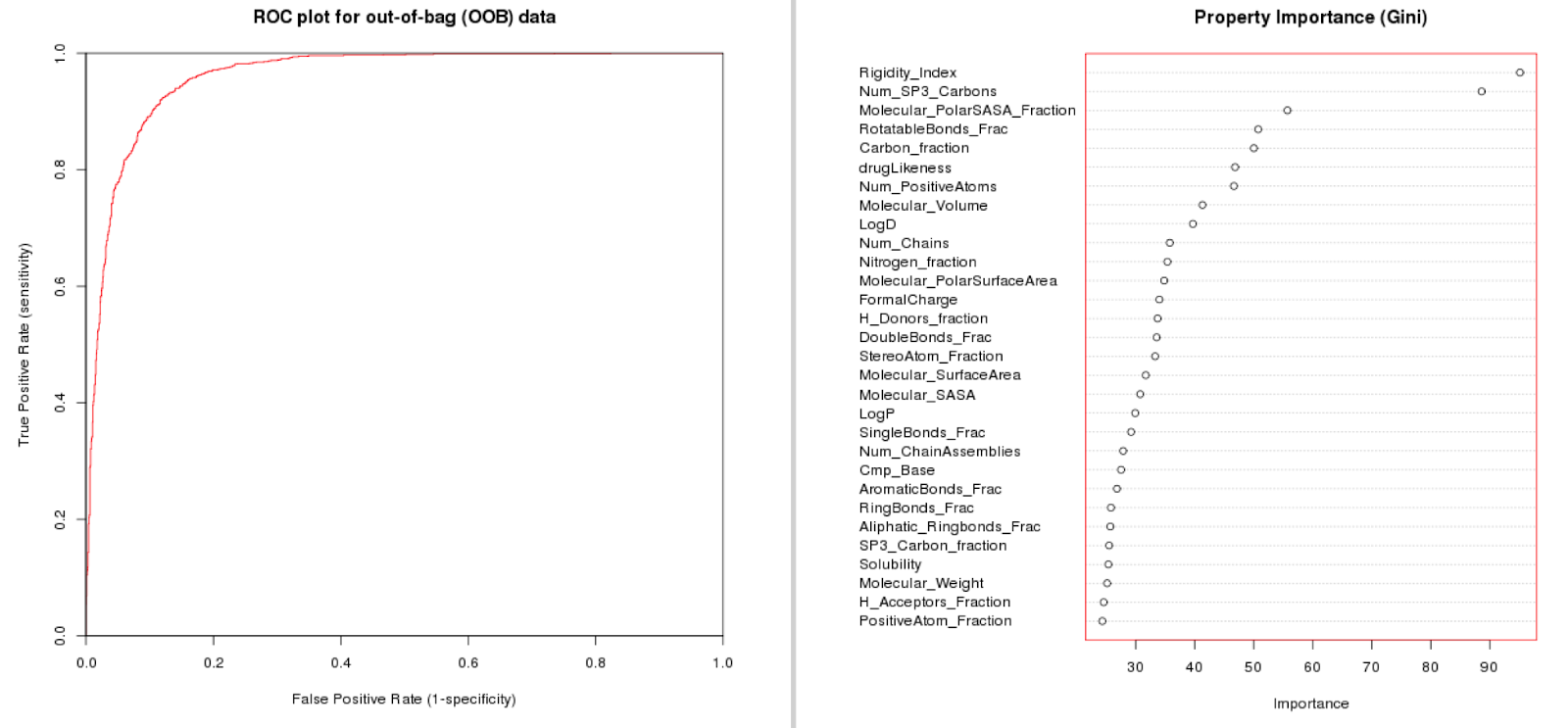
Built on the dataset we have created allosteric classifier models that can predict if an interaction is likely allosteric or not. We have tried this on the full dataset, but also on lower levels (restricting the data to e.g. Class A GPCRs). We find that we can train predictive models that gain in quality if we have a more concise dataset (eliminating some of the inter-target variation). In the paper we provide case studies on HIV Reverse Transcriptase, the adenosine receptors (family), and protein Kinase B. Here the model performance for class A GPCRs (full L2 tgt class) is shown. Note that rigidity, number of sp3 carbons, Polar Solvent Accessible Surface (normalized), and rotatable bonds fraction are most important for model fit.
All data is ChEMBL and hence can be freely downloaded and used. Please let us know if you find any errors or misclassifications as we will correct them (crowd curation).
Anna, jpo, and Gerard
%T Chemical, Target, and Bioactive Properties of Allosteric Modulation
%A G.J.P. van Westen
%A A. Gaulton
%A J.P. Overington
%J PLoS. Comput. Biol.
%D 2014
%V 10
%O doi:10.1371/journal.pcbi.1003559