-
The great US patent spike on SureChEMBL
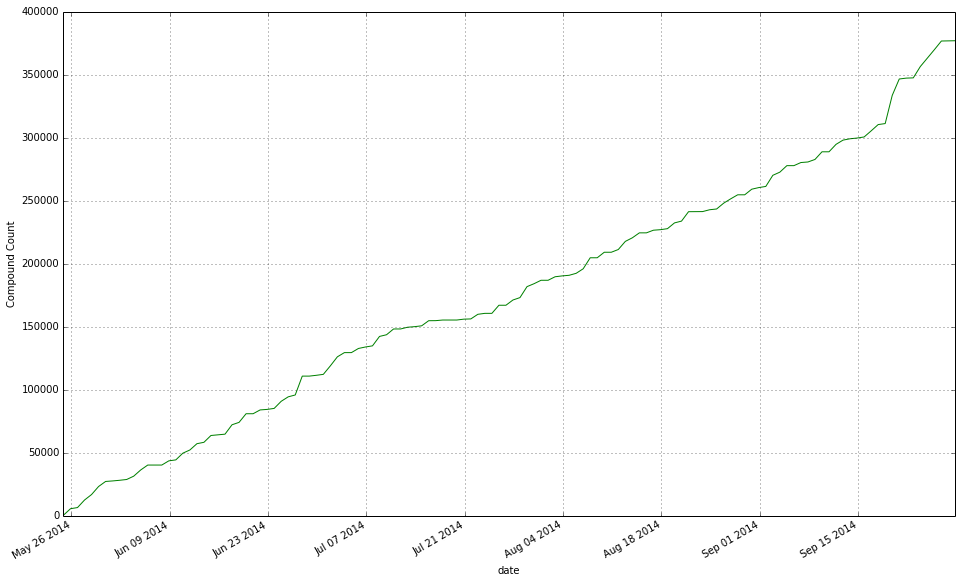
Apparently, there was a huge spike of new granted US patents released by the USPTO a few days ago. The reason?In March 2013, US patent law changed. The ‘first to invent’ became ‘first inventor to file’ for patent protection purposes (see more on this here). As a result, a lot of people rushed to submit applications just before the change. Fast forward 18 months later (last week), a huge spike in USPTO granted patents is observed.Did SureChEMBL pick that up? See below the cumulative count plot of new patent documents:And the corresponding compound count extracted from these patents:For more information on SureChEMBL, see our previous posts.George -
SureChEMBL Available Now
Followers of the ChEMBL group's activities and this blog will be aware of our involvement in the migration of the previously commercially available SureChem chemistry patent system, to a new, free-for-all system, known as SureChEMBL. Today we are very pleased to announce that the migration process is complete and the SureChEMBL website is now online.SureChEMBL provides the research community with the ability to search the patent literature using Lucene-based keyword queries and, much more importantly, chemistry-based queries. If you are not familiar with SureChEMBL, we recommend you review the content of these earlier blogposts here and here. SureChEMBL is a live system, which is continuously extracting chemical entities from the patent literature. The time it takes for a new chemical in the patent literature to become searchable in the SureChEMBL system is 1-2 days (WO patents can sometimes take a bit longer due to an additional reprocessing step). At time of writing this blogpost the number of unique compounds in SureChEMBL is 15,760,514, which have been extracted from 12,949,021 patents.To get started using SureChEMBL, head over to the homepage, where you will be presented with a range of search methods and filters. The image below provides a brief overview of the search functionality offered by the system:
To provide an example of how to use the SureChEMBL website, let's assume you are interested in patents which contained structures similar (or identical) to Sildenafil in the claims section of the document and also mention the term PDE5 anywhere in the document. To run this search, go to the SureChEMBL homepage and carry out the following actions:- Enter the term 'PDE5' in the search text box
- Sketch in the structure of Sildenafil (or use the name look-up function)
- Change the search type to similarity (>85%)
- Click the 'Claims' checkbox in the document filter section and
- Hit 'Search' button
After clicking 'Search', you will be presented with a page which contains all compounds that match your search criteria:
From the compound results page above you then have the choice of either exporting the chemistry (all the compounds returned by the search) or viewing the patents associated with 1 more of the selected compounds. For the selected compounds in this search, the associated patents (sorted by descending publication date) are :
From the patent document results page, you are able to export chemistry from all documents on display, view patent family information and view the chemistry-annotated, full text document. The claims section of the first patent (US-20140255433-A1) includes references to both sildenafil and PDE5:
The aim of this blogpost is to introduce the SureChEMBL system and not to provide a comprehensive review of all the functionality the system offers. This will be covered in future training sessions and webinars, which will be announced on this blog in the near future.We would like to thank the people over at Digital Science, who were responsible for building the original SureChem system and supported its migration over to EMBL-EBI. In particular, we would like to thank Nicko Goncharoff, James Siddle and Richard Koks.The system runs on the cloud - specifically on Amazon Web Services, a stable, secure and highly scalable way to deploy web applications. We need to keep a close eye on performance and patterns of usage over the coming weeks, to get an idea of how many servers, etc, we need for full deployment. In particular, we will throttle scripted access, so please get in touch if you want to try anything like this, so you are not frustrated by slow performance, and we will try and accommodate your use case. There is also a download link on the homepage, so please explore this if you are interested.
We have an exciting roadmap for the future development of SureChEMBL, bt if you have any priority requests, mail them to surechembl-help (at) ebi.ac.uk.
If you experience any issues with the system, or have any questions please get in touch. -
Another Confusion in the Literature - Trust but Verify
Another kinase inhibitor mystery solved! I (jpo) feel like I'm turning into a rotund Hercule Poirot, except English and with a huge fat wirey 'tache.
The literature has quite a few references to an Trk inhibitor in phase 2 trials from Cephalon - CEP-2583; just do a google search with something like "CEP-2583 clinical trial" and you'll come across a number of references to this, usually in the peer-reviewed literature. However, this is really the only place is occurs, try clinicaltrials.gov, Cephalon regulatory filings, Teva pipeline, nothing.
However, this small letter to Journal of Urology (DOI:s10.1097/01.ju.0000138215.70709.9c) solves the mystery clinical candidate - it was a typo, it was meant to refer to CEP-2563/CEP-701 (a prodrug/drug pair), and so a simple transcription error (8 for 6), and then repetition across other reviews (who I guess trusted the primary source) gave a footprint to the existence of a clinically interesting asset, that never was.
What to do with these sort of errors in the literature, the are too minor for journals to ever worry about fixing, but arguably they should if bought to their attention. But of course, post-publication peer review should also fix this. So I'll make some comments on pubmedcommons when I have some time. It was good of Cephalon to correct the record in this way.
jpo, Bissan, and Krister -
Papers: Literature text mining and extensions to UniChem
Two new papers from the group have just been published, both in Journal of Chemoinformatics - and of course both Open Access.The first deals with some extensions to UniChem to allow far more flexible searches. The abstract is:UniChem is a low-maintenance, fast and freely available compound identifier mapping service, recently made available on the Internet. Until now, the criterion of molecular equivalence within UniChem has been on the basis of complete identity between Standard InChIs. However, a limitation of this approach is that stereoisomers, isotopes and salts of otherwise identical molecules are not considered as related. Here, we describe how we have exploited the layered structural representation of the Standard InChI to create new functionality within UniChem that integrates these related molecular forms. The service, called ‘Connectivity Search’ allows molecules to be first matched on the basis of complete identity between the connectivity layer of their corresponding Standard InChIs, and the remaining layers then compared to highlight stereochemical and isotopic differences. Parsing of Standard InChI sub-layers permits mixtures and salts to also be included in this integration process. Implementation of these enhancements required simple modifications to the schema, loader and web application, but none of which have changed the original UniChem functionality or services. The scope of queries may be varied using a variety of easily configurable options, and the output is annotated to assist the user to filter, sort and understand the difference between query and retrieved structures. A RESTful web service output may be easily processed programmatically to allow developers to present the data in whatever form they believe their users will require, or to define their own level of molecular equivalence for their resource, albeit within the constraint of identical connectivity.The second deals with using text mining approaches to find papers that look like they could be abstracted into ChEMBL - that is they contain keywords enriched in medicinal chemistry and compound structure concepts. The abstract for this paper is:
The large increase in the number of scientific publications has fuelled a need for semi- and fully automated text mining approaches in order to assist in the triage process, both for individual scientists and also for larger-scale data extraction and curation into public databases. Here, we introduce a document classifier, which is able to successfully distinguish between publications that are ‘ChEMBL-like’ (i.e. related to small molecule drug discovery and likely to contain quantitative bioactivity data) and those that are not. The unprecedented size of the medicinal chemistry literature collection, coupled with the advantage of manual curation and mapping to chemistry and biology make the ChEMBL corpus a unique resource for text mining.The method has been implemented as a data protocol/workflow for both Pipeline Pilot (version 8.5) and KNIME (version 2.9) respectively. Both workflows and models are freely available at: ftp://ftp.ebi.ac.uk/pub/databases/chembl/text-mining. These can be readily modified to include additional keyword constraints to further focus searches.Large-scale machine learning document classification was shown to be very robust and flexible for this particular application, as illustrated in four distinct text-mining-based use cases. The models are readily available on two data workflow platforms, which we believe will allow the majority of the scientific community to apply them to their own data.%T UniChem: extension of InChI-based compound mapping to salt, connectivity and stereochemistry layers %A J Chambers %A M Davies %A A Gaulton %A G Papadatos %A A Hersey %A JP Overington %J Journal of Cheminformatics %D 2014 %V 6:43 %O doi:10.1186/s13321-014-0043-5 %O http://www.jcheminf.com/content/6/1/43 %T A document classifier for medicinal chemistry publications trained on the ChEMBL corpus %A G Papadatos %A GJP van Westen %A S Croset %A R Santos %A S Trubian %A JP Overington %J Journal of Cheminformatics %D 2014 %V 6:40 %O doi:10.1186/s13321-014-0040-8 %O http://www.jcheminf.com/content/6/1/40
-
Structure Confirmation, Reproducibility of Research, and Literature Abstraction - Trust But Verify
The identity of compounds in the literature is often uncertain - with high profile cases of incorrect compounds being used in experiments - with subsequent difficulties in reconciling conflicting experiments or repeating results.
One example is the kinase inhibitor Bosutinib (SKI-606) - where several compound vendors shipped a similar isomer of the actual drug. It will be unclear which data comes from the bona fide Bosutinib and the compound sold as Bosutinib by multiple vendors. More details on this case are in this post by Derek Lowe, and there is a definitive structural biology paper here. So this probably clouds a lot of the reported Bosutinib bioactivity data that's out there. Here the difference is in the differential substitution pattern on an aromatic ring (2,4-dichloro-5-methoxy vs 3,5-dichloro-4-methoxy), the mass is the same for these isomers.
A second example is this case (a clinical stage compound called TIC10) where there was a mix-up in the structure assigned to a compound in a patent, and since the substance had novel and potentially useful bioactivity and the annotated structure in wrong, led to all sorts of complicated IP issues and shenanigans. Since this compound was also in a widely distributed and profiled NCI set, then the literature ends up with difficult to deconvolute data.
There are probably many of these examples - and here are two more, both connected to clinical development stage protein kinase inhibitors from Exelixis.
The first is XL-765 also known as SAR-245409 and by the INN Voxtalisib - it is a PI3K inhibitor. Compound vendors have been selling 'XL-765' for some time, but the structure sold is (typically) not the correct XL-765 structure.
XL-765 is known to be this structure - multiple independent authoritative source point to this identity. It's a quite simple, small molecule. The InChI key is RGHYDLZMTYDBDT-UHFFFAOYSA-N.
Several vendors have been selling a different structure as 'XL-765/SAR-245409'; as you will immediately note, it is a very different structure - not a positional or stereo-isomer, it will have very different mass, etc.
Some purchasers of this compound report peer-reviewed in vivo data. To be clear, this isn't just one company, it is all/most vendors of XL-765.
So there are a number of things that could have happened here:
i) The correct structure was supplied, but the structure on the label/on the website didn't match the real physical structure - i.e. a mislabelling.
ii) The incorrect structure was supplied, and the structure on the label/website was what was in the bottle. In our studies (at FIMM) the vendor supplied XL-765 has not behaved as a PI3K inhibitor, so the latter is more likely.
However, the incorrect XL-765 structure has propagated into public chemistry databases, and literature data for XL-765 is probably now largely suspect (with the exception of that from Exelixis themselves (and their collaborators)).
Here's the second new example - XL-147 (aka SAR-245408 and Pilaralisib) - again a PI3K inhibitor. The correct structure is QINPEPAQOBZPOF-UHFFFAOYSA-N.
Many vendors were selling XL-147 as the following compound, here there is a lot more similarity between the actual and sold compound, but it's still different, and any biology on this, although it may well be active in it's own right, is not the same as XL-147.
tl;dr Be careful if you are interested in purchasing or analysing activities and properties of Bosutinib, Pilaralisib &Voxtalisib. Be careful in loading names from Vendor catalogs, and try to use more definitive authorities for compound synonyms. Be careful with relying on things from any public databases.
Krister and I would like to thank Willie Yuan at Chemietek for helping to clarifying aspects of the confusing history for this compound.
jpo and Krister -
We're hiring! Web developer for NIH Illuminating the Druggable Genome (IDG) project
We got a prize today, so we are happy. What better way to celebrate, than to recruit someone new for the group. We have a position available for a developer to support web service development and integration for the Knowledge Management Centre part of the recently announced NIH Illuminating the Druggable Genome project, see this link for details of the job.
Closing deadline for applications is 12th October 2014. -
SureChEMBL Update 1
As announced in the previous SureChEMBL blogpost, the temporary holding page is now in place. So when users visit https://www.surechem.com (or https://open.surechem.com), you will be redirected to https://www.surechembl.org.
For updates on the release of the new SureChEMBL site, please keep an eye on the ChEMBL-og. -
SureChEMBL Coming Very Soon
In the coming weeks we will be very pleased to announce the release of the new SureChEMBL website. Since the beginning of the year, we have been working hard with the folks over at Digital Science, along with all the content and software providers to get the system setup and running on our own Amazon Web Service controlled environment. As we approach the final stages of the transition, we will need to temporarily halt access to the original SureChem site. The reason for this minor disruption is to allow us to complete the testing of the additional functionality we have added to the SureChEMBL user interface.
We will use ChEMBL-og as the primary route of communicating with users, so if you want to be kept up to date, bookmark the site. We will also make ad hoc tweets about SureChEMBL on @johnpoverington, @georgeisyourman, @surechembl and @chembl.
SureChEMBL User Interface
Users familiar with the previous SureChem UI will find a lot in common with the new SureChEMBL UI. A summary of the changes and new features we have added to the SureChEMBL UI are provided below:- A user account is no longer required to access the system
- All users will have access to ‘Pro’ account features, which include chemistry exports, PDF downloads and enhanced search filters
- UniChem has been integrated and provides dynamic cross references to external chemical resources
- The new SCHEMBL identifier is used throughout the interface.
- Updated compound sketchers (Latest Marvin JS and JSME)
- Rebranding of headers and footers and removing old SureChem references
SCHEMBL Identifier
In line with ChEMBL IDs, all compounds in SureChEMBL have been given SCHEMBL identifiers. For example, SCHEMBL1353 corresponds to 2-(acetyloxy)benzoic acid, aka aspirin. The identifier can be used to access the SureChEMBL compound page and will be included in all SureChEMBL downloads.
SureChEMBL Data Content
The SureChEMBL pipeline has been running daily throughout the summer and has now processed and extracted an additional ~400,000 novel compounds from patents since SureChem’s pipeline freeze. At the time of writing (16:22 22/08/14), the SureChEMBL counts are:- Total number of compounds 15,668,22
- Total number of annotated patents 12,888,125
The rate of novel compounds and annotated patents is truly staggering: There are approximately 80,000 compounds extracted from 50,000 patents that are added to the system every month. Moreover, the latency for a new patent document from its application date to becoming searchable in the system is only between 2 and 7 days, in most cases.
SureChEMBL and UniChem
The complete SureChEMBL structure repository has been added to UniChem (src_id=15) and consists of 15.2M unique structures mapped to their SCHEMBL IDs. SureChEMBL updates will be added to UniChem on a weekly basis, so that UniChem will be up to date with novel patent chemistry.
SureChEMBL Data Access
Besides availability in UniChem, the complete SureChEMBL structure repository is provided as SD and tsv file in our ftp site:It has to be emphasised here that this is the raw compound feed as extracted automatically from text and images and is provided without any further filtering or manual curation. This feed contains fragments, radicals, atoms with wrong valencies, polymers and other oddities but if you are the sort of person who wants to use this raw data, you will know what and how to filter things you don't like out.
The chemical registry rules between SureChEMBL and ChEMBL have not been fully aligned yet - they use fundamentally different toolkits - so there are sometimes multiple SCHEMBL ids for the same InChI - if you know this is an issue, you will know how to fix it for your local purposes if you download the data.
Initially, the SureChEMBL files on the ftp site will be updated on a quarterly basis.
SureChEMBL Future Plans
Going forward we have many plans related to SureChEMBL, some of which are linked to our involvement in the Open PHACTS project. Our current plans include:- Extraction of biological entities from the patent literature
- SureChEMBL API release
- Updated workflow tool integration (e.g. KNIME and Pipeline Pilot)
You will hear more about these plans over the coming year, but our top priority now is to deliver the new SureChEMBL user interface.
If you have any questions about the new SureChEMBL system and data please get in touch